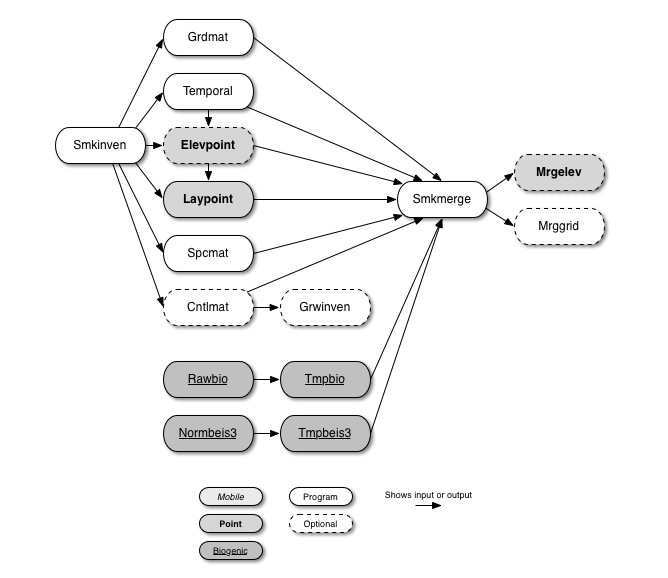
Figure 6.1, “SMOKE core programs” shows the SMOKE core programs and how they relate to each other. We define the core programs as those needed for the actual emissions processing; we exclude QA and utility programs such as Smkreport (these programs are discussed in Chapter 7, SMOKE Quality Assurance and Chapter 5, SMOKE Utility Tools). The lines between the boxes represent one or more files passing between the programs, and show the dependencies among the programs. Note that we have omitted some redundant file dependencies to make the diagram easier to understand. Unless otherwise noted, the SMOKE core programs are applied for the anthropogenic source categories: area-, mobile-, and point-source processing (biogenic processing follows a separate path from the other source categories). When a program that applies to multiple source categories is used, it can be run for only one source category at a time, with the exception of Smkmerge and Mrggrid.
The Smkinven program, at left in Figure 6.1, “SMOKE core programs”, is responsible for importing the stationary area/non-point, nonroad, on-road mobile, and point source inventory emissions data. For mobile sources, Smkinven can also import activity data in the form of VMT and vehicle speed for use in generating emission factors. For point sources, Smkinven can also import day- and hour-specific data. The output from Smkinven is used as input to nearly every other core SMOKE program.
Also at left, the Rawbio program imports the county or gridded land use data and computes normalized (time-independent) biogenic emissions.
As an alternative to BEIS2 processing using Rawbio, SMOKE is capable of using the BEIS3 model. In this type of processing, Normbeis3 creates gridded, normalized biogenic emissions from land use and biogenic emissions factors.
Grdmat creates the gridding matrix for the anthropogenic source categories.
The Temporal program is used to create an hourly emissions file for the anthropogenic source categories. It can read in day-specific and hour-specific data, and merge this with estimated daily and hourly data created from the annual emissions data using temporal profiles.
Elevpoint preprocesses the selected PinG and major point sources. Elevpoint is not used when no PinG or major point sources need to be defined. In this case, SMOKE can create elevated emissions for all point sources, so there is no need to specifically indicate the major point sources.
Laypoint computes the plume rise for all point sources based on the meteorology data.
Spcmat creates the speciation matrices (both mass and molar) for the anthropogenic source categories. It uses the user-selected chemical mechanism.
Cntlmat creates the growth matrices, multiplicative control matrices, and reactivity control matrices for the anthropogenic source categories. This program is not used for base-year emissions without controls.
Grwinven is used to grow the emissions to past or future years using the growth matrix created by Cntlmat and the imported inventory data from Smkinven. It writes both SMOKE inventory format (I/O API NetCDF) and IDA format. The output from Grwinven is used in place of the original output from Smkinven when processing past or future years.
Tmpbio applies meteorology adjustments to the gridded, normalized biogenic emissions from Rawbio. It also applies the speciation factors needed for the user-selected chemical mechanism to create gridded, hourly, model-species biogenic emission files for use in Smkmerge or Mrggrid. The Rawbio and Tmpbio programs taken together are the equivalent of SMOKE-BEIS2.
Tmpbeis3 applies meteorology adjustments to the normalized emissions created by Normbeis3. Tmpbeis3 also applies the speciation profiles needed for the user-selected chemical mechanism to create gridded, hourly, model-species biogenic emissions data for use in Smkmerge and Mrggrid. The Normbeis3 and Tmpbeis3 programs taken together are the equivalent of SMOKE-BEIS3.
Smkmerge is used to combine all emissions and matrices to create the gridded, hourly, model-species emissions needed for an AQM. It can merge for one source category at a time or all source categories at once, and it can read in the model-ready biogenic emissions to merge with the anthropogenic source categories. It can merge the matrices with the inventory data output from Smkinven or the hourly emissions from Temporal, and it can optionally merge the speciation, gridding, or control matrices, or any combination. It also writes state and county emissions totals.
Movesmrg is only used for mobile sources to compute hourly emissions using the hourly VMT from Temporal, the MOVES emission rates lookup tables (i.e., RPD, RPV and RPP), the gridded/averaged temperatures meteorology data and more. If VMT are not used, then the emissions from Smkinven are combined with the temporal factors, and the emission factors are not used at all.
In the remaining sections of this chapter, we provide detailed program descriptions, in alphabetic order.
Mrgelev is used to combine ASCII elevated files created by Smkmerge. These files contain point-source data needed by AQMs including CAMx, REMSAD, and UAM models.
Mrggrid is used to combine gridded emission data files, which can be speciated or non-speciated, and hourly or time-independent. It can combine a 3-D point source file with any number of 2-D files from other source categories. This program is optional and provides a convenient way to merge model-ready output files outside of Smkmerge.